Simulation benchmarks¶
New in version 11.2.0.
See also
As of version 11.2.0, ixpeobssim provides support for simulation benchmarks, or advanced analysis pipelines that are specifically designed to generate a large number of different realizations of the same observation to gauge the statistical properties of a given measurement (e.g., the posterior distribution of the fit parameters).
If you are familiar with the concept of an ixpeobssim analysis pipeline, this is no different, and you can peek at one of the examples, e.g. [github]/ixpeobssim/benchmarks/toy_polconst.py, to take inspiration for your own first benchmark. The rest of this section provides some additional, specific information.
Generating ensambles¶
The ixpeobssim.core.pipeline module provides a mean to generate
a simulation ensamble through the
ixpeobssim.core.pipeline.generate_ensamble() method. The following
minimal snippet, for instance, will generate 1000 different realizations of
a 50 ks observation with the toy_pollin model.
import ixpeobssim.core.pipeline as pipeline
pipeline.bootstrap_pipeline('toy_pollin')
pipeline.generate_ensamble(size=1000., duration=50000.)
One (non-obvious) feature of this method is that the processing of the output event lists is embedded in the generation and, by default, the method cleans up after itself by deleting from disk all the (potentially very large) event files, only leaving the processing products (e.g., binned files and/or fit output). This is done because if you are generating a 1000 realization of an observation with many events, you might easily run out of disk space before you realize it—and, after all, once the necessary processing is done you don’t need the event files anymore.
By default the pipeline will call
ixpeobssim.core.pipeline.standard_ensamble_processing() under the hood
(which, in turn, will create Stokes spectra and modulation cubes), but you can
override this behavior and use your own custom processing function by just doing
something along the lines of:
import ixpeobssim.core.pipeline as pipeline
def process(file_list):
"""Implement your own processing function here!
"""
pipeline.xpbin(*file_list, algorithm='PHA1')
pipeline.bootstrap_pipeline('toy_pollin')
pipeline.generate_ensamble(size=1000., duration=50000., processing_function=process)
Post-processing ensambles¶
Once you have generated and processed all your simulation, you typically want to post-process the binned files, extract the relevant parameters, and store the results for later use (e.g., statistical analysis and/or plotting). As an example, you might want to fit the Stokes spectra with XSPEC with the proper spectro-polarimetric model and store the fit results in a table.
ixpeobssim provides a few facilities for doing just this, including:
a mechanism for easily retrieving the path to the relevant data product, modeled on that of the simple pipelines;
a data table class to store and write to FITS the results of the post-processing.
You should look at [github]/ixpeobssim/benchmarks/toy_polconst.py for a real-life example.
An illustrative use case¶
The [github]/ixpeobssim/benchmarks/toy_polconst.py benchmark is possibly the easiest toy setup resembling a real observation.
The source model is [github]/ixpeobssim/config/toy_point_source.py is a point source with a power-law spectrum and a constant polarization degree and angle. The benchmark simulates 1000, 50 ks-long observations (mind this is a bright source, and a 50 ks observation time accounts for some 10,000,000 events—so this is a high-statistics benchmark by all metrics).
At the post-processing stage we feed the I, Q and U Stokes spectra into XSPEC
and fit them with a powerlaw * polconst spectro-polarimetric model.
Below are the pulls
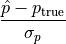
for all the four fit parameters—the spectral index and normalization, and the polarization degree and angle. The fact that they are all reasonably distributed as standard normal variables, with zero mean and unit standard deviation, is a sensible sanity check for the entire analysis chain.
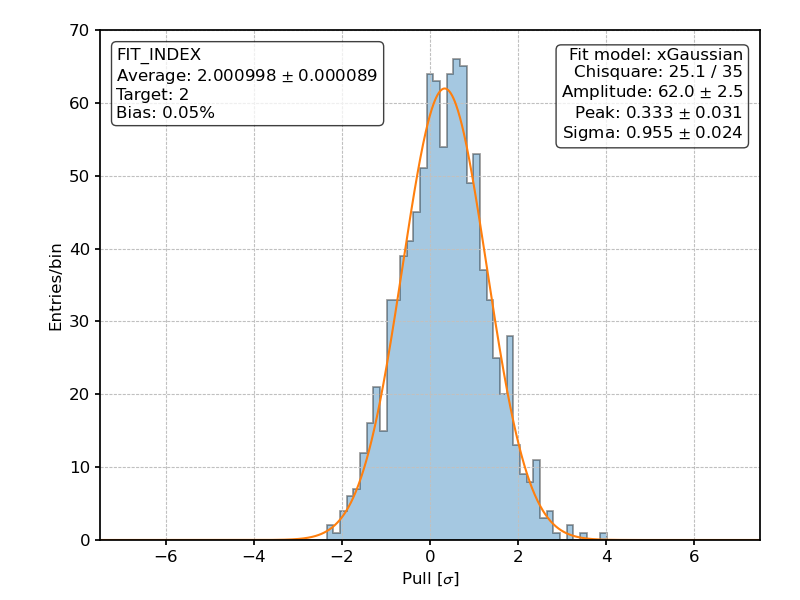
Posterior distribution of the pulls for the fitted spectral index over 1000 realization of the [github]/ixpeobssim/benchmarks/toy_polconst.py benchmark.¶
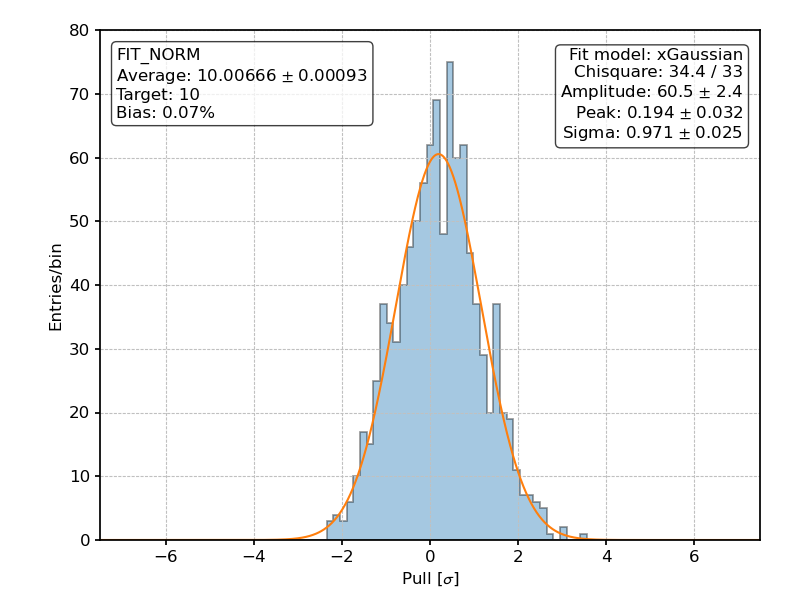
Posterior distribution of the pulls for the fitted spectral normalization over 1000 realization of the [github]/ixpeobssim/benchmarks/toy_polconst.py benchmark.¶
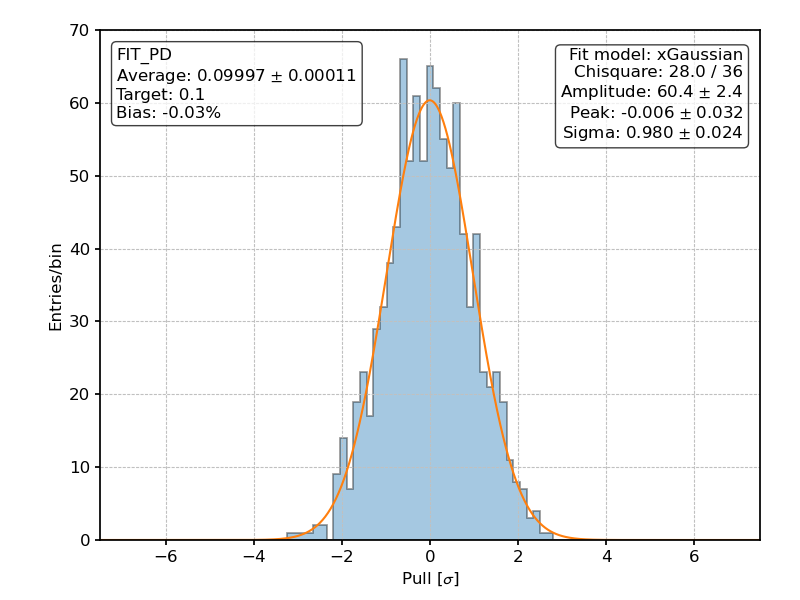
Posterior distribution of the pulls for the fitted polarization degree over 1000 realization of the [github]/ixpeobssim/benchmarks/toy_polconst.py benchmark.¶
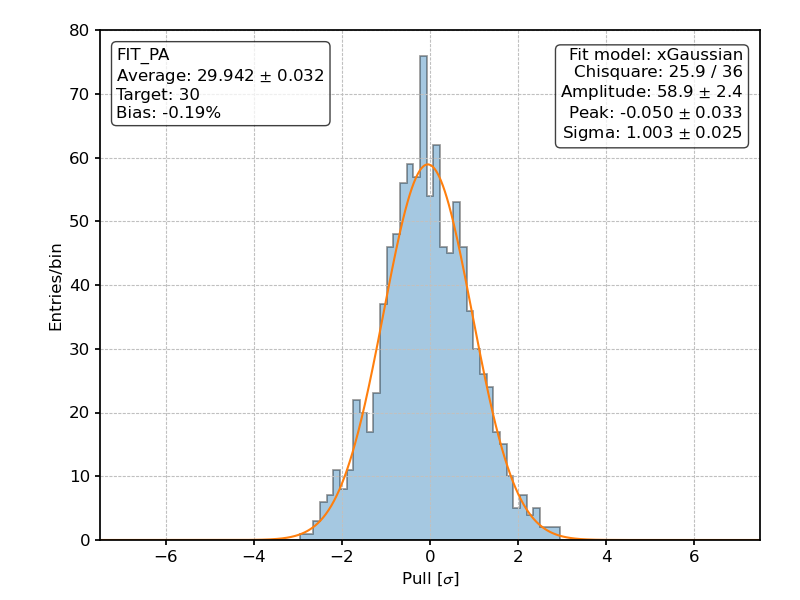
Posterior distribution of the pulls for the fitted polarization angle over 1000 realization of the [github]/ixpeobssim/benchmarks/toy_polconst.py benchmark.¶
